
Discovery of novel carbohydrate-active enzymes through the rational exploration of the protein sequences space | PNAS
Bioinformatics as a tool for assessing the quality of sub-cellular proteomic strategies and inferring functions of proteins: plant cell wall proteomics as a test case - Document - Gale Academic OneFile

The Ruminococcus bromii amylosome protein Sas6 binds single and double helical α-glucan structures in starch | bioRxiv

Reaction Mechanisms in Carbohydrate-Active Enzymes: Glycoside Hydrolases and Glycosyltransferases. Insights from ab Initio Quantum Mechanics/Molecular Mechanics Dynamic Simulations | Journal of the American Chemical Society
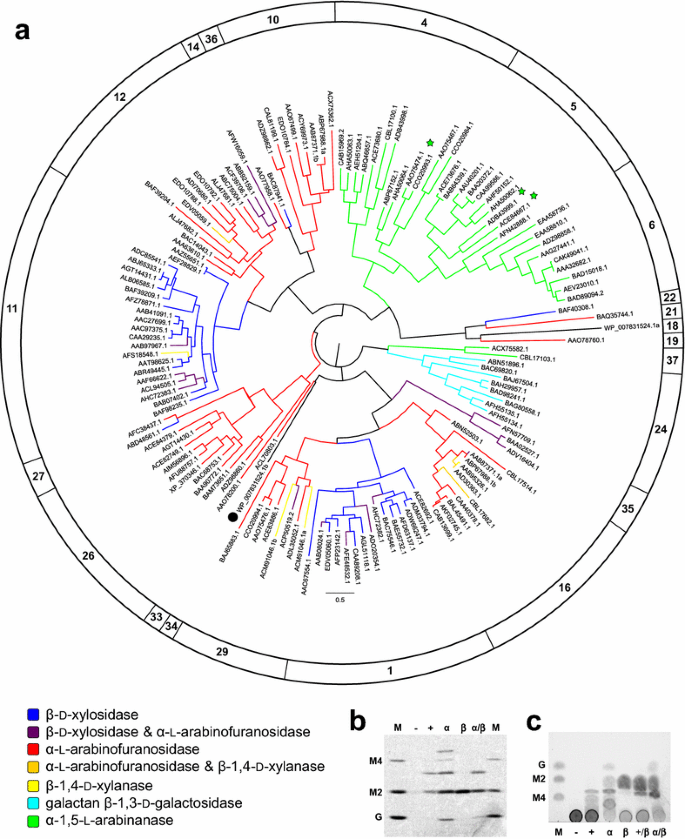
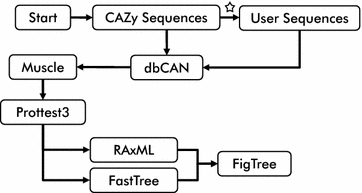
SACCHARIS: an aed pipeline to streamline discovery of carbohydrate active enzyme activities within polyspecific families and de novo sequence datasets | Biotechnology for Biofuels and Bioproducts | Full Text

Chitin-Active Lytic Polysaccharide Monooxygenases Are Rare in Cellulomonas Species | Applied and Environmental Microbiology

An introduction On behalf of the ”CAZypedia Consortium” Harry Brumer, Ph.D. Professor The Michael Smith Laboratories, UBC and School of Biotechnology, - ppt download
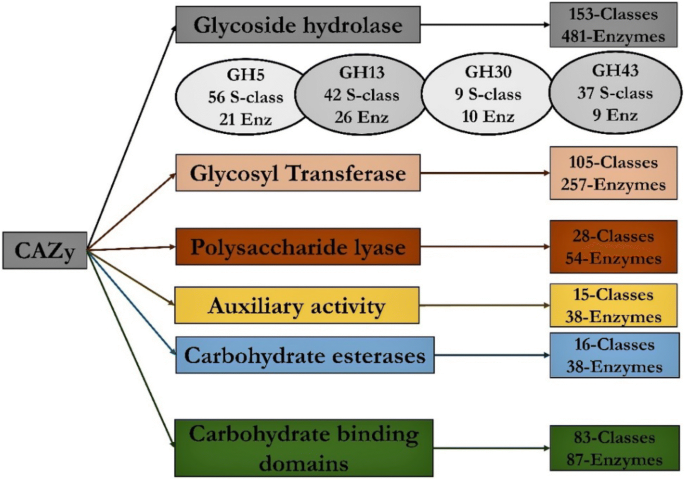
CAZymes-based ranking of fungi (CBRF): an interactive web database for identifying fungi with extrinsic plant biomass degrading abilities | Bioresources and Bioprocessing | Full Text

PDF) The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics | Thomas Bernard - Academia.edu

Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes – topic of research paper in Biological sciences. Download scholarly article PDF and read for free on CyberLeninka
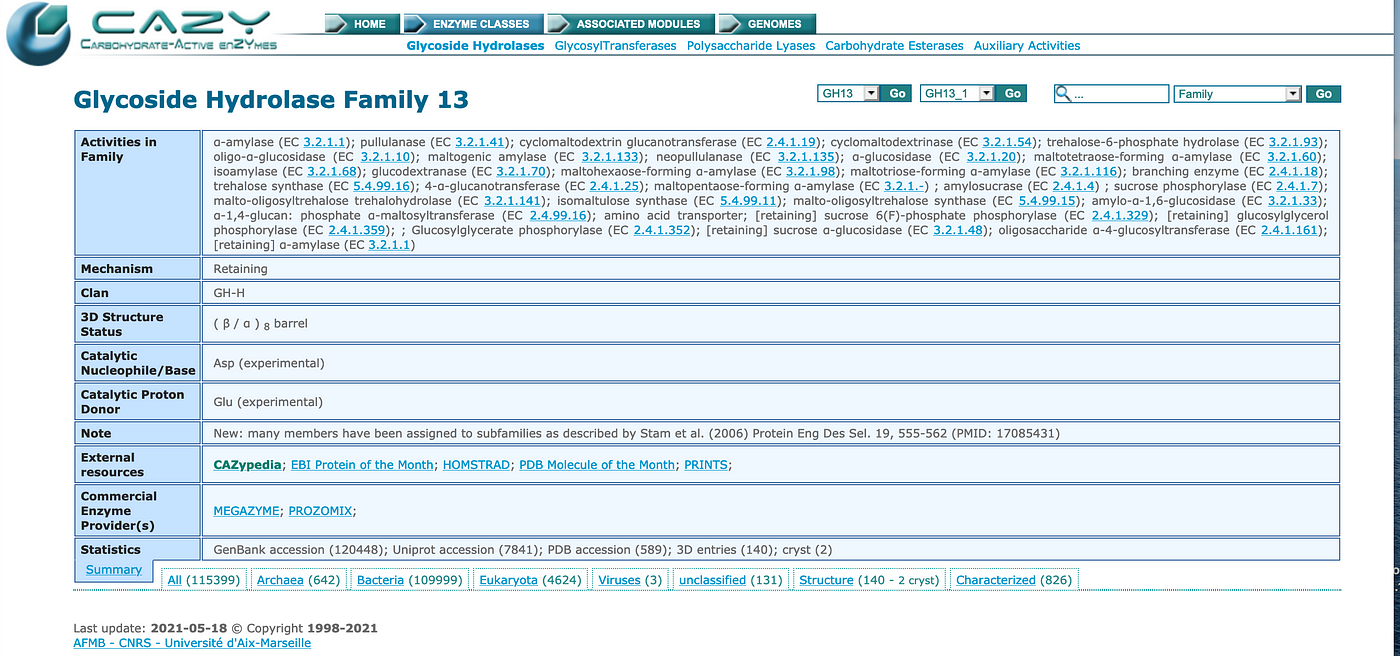
Graph Database, GraphQL and Machine Learning for Carbohydrate-Active Enzymes | by Sixing Huang | Towards Data Science

A carbohydrate-active enzyme (CAZy) profile links successful metabolic specialization of Prevotella to its abundance in gut microbiota | Scientific Reports
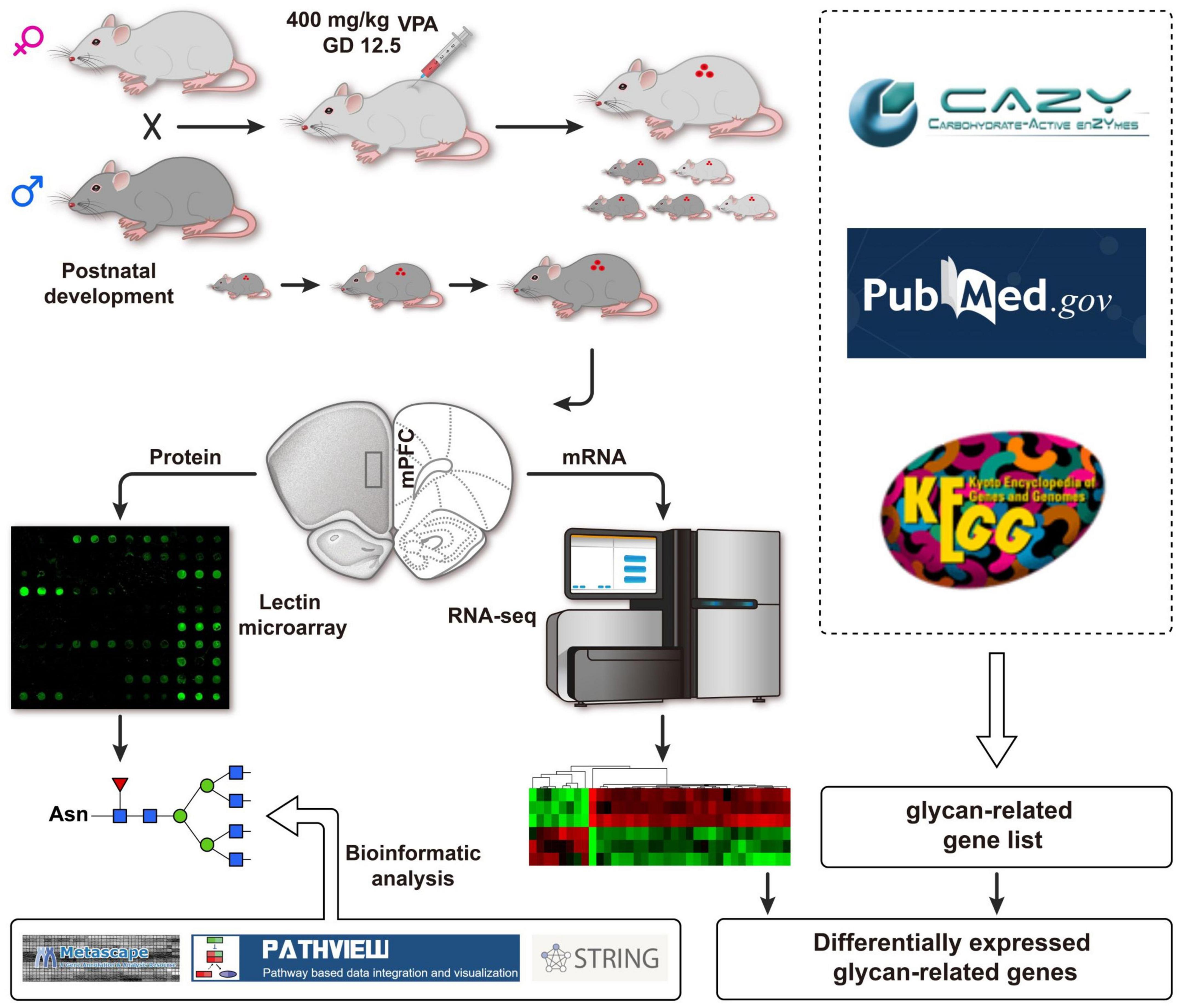
Frontiers | Altered expression of glycan patterns and glycan-related genes in the medial prefrontal cortex of the valproic acid rat model of autism
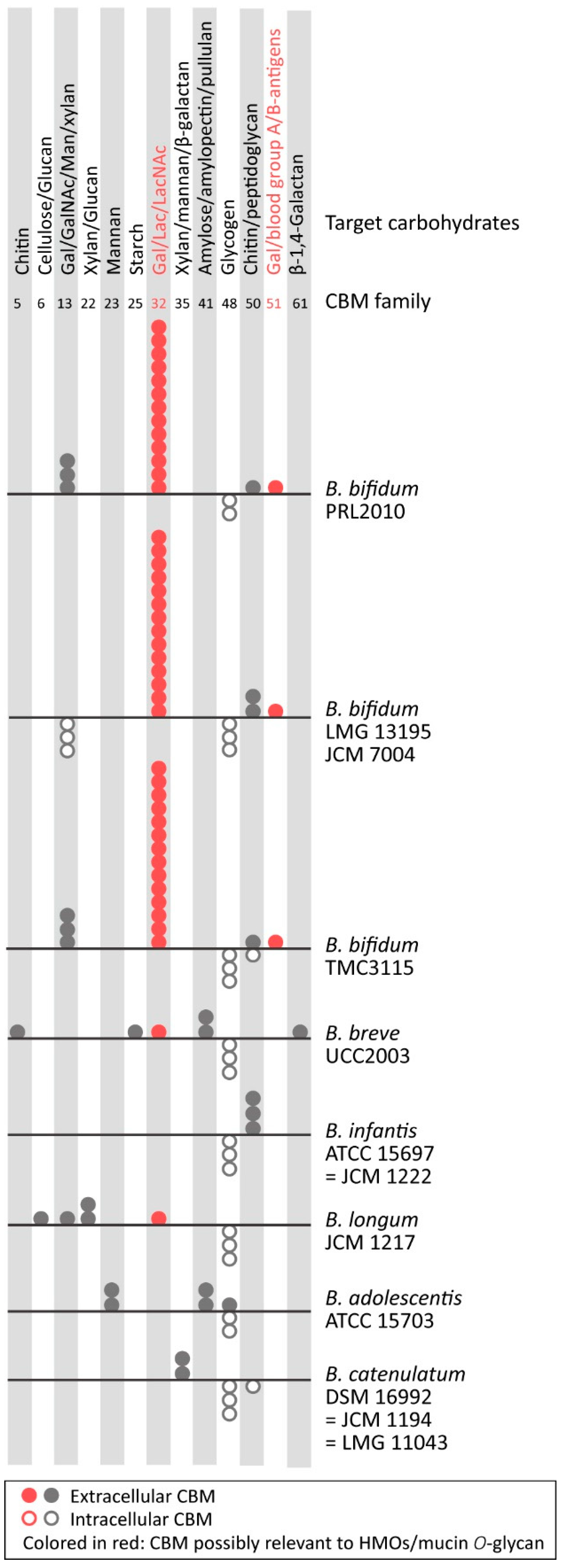

Microorganisms | Free Full-Text | Enzymatic Adaptation of Bifidobacterium bifidum to Host Glycans, Viewed from Glycoside Hydrolyases and Carbohydrate-Binding Modules
![PDF] The carbohydrate-active enzymes database (CAZy) in 2013 by Vincent Lombard, Hemalatha Golaconda Ramulu, Elodie Drula, Pedro M. Coutinho, Bernard Henrissat · 10.1093/nar/gkt1178 · OA.mg PDF] The carbohydrate-active enzymes database (CAZy) in 2013 by Vincent Lombard, Hemalatha Golaconda Ramulu, Elodie Drula, Pedro M. Coutinho, Bernard Henrissat · 10.1093/nar/gkt1178 · OA.mg](https://og.oa.mg/The%20carbohydrate-active%20enzymes%20database%20(CAZy)%20in%202013.png?author=%20Vincent%20Lombard,%20Hemalatha%20Golaconda%20Ramulu,%20Elodie%20Drula,%20Pedro%20M.%20Coutinho,%20Bernard%20Henrissat)
PDF] The carbohydrate-active enzymes database (CAZy) in 2013 by Vincent Lombard, Hemalatha Golaconda Ramulu, Elodie Drula, Pedro M. Coutinho, Bernard Henrissat · 10.1093/nar/gkt1178 · OA.mg
The carbohydrate-active enzymes database (CAZy) in 2013. : Lombard, Vincent : Free Download, Borrow, and Streaming : Internet Archive

Function of CAZymes encoded by highly abundant genes in rhizosphere microbiome of Moringa oleifera - ScienceDirect




![PDF] The carbohydrate-active enzymes database (CAZy) in 2013 | Semantic Scholar PDF] The carbohydrate-active enzymes database (CAZy) in 2013 | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/0e5bccdedb82fbafece8ca71d64b16ff05ec9145/5-Table2-1.png)


